Translation Vs. Transcription: Similarities And Differences
Maybe your like
Every protein in your body — every enzyme, receptor, antibody, and structural fiber — began as a gene encoded in DNA. But DNA never leaves the nucleus. The cell solves this problem with a two-step relay: transcription copies the gene into a portable RNA message, and translation reads that message to assemble a protein. Together, these processes form the core of the central dogma of molecular biology (DNA → RNA → protein), and they appear on the AP® Biology exam in every format: diagram interpretation, sequence-tracing MCQs, and FRQs that ask you to predict what happens when a specific step is disrupted.
This article walks through both processes stage by stage, with the molecular details, comparison tables, and exam connections you need for test day.
What We Review
- What is Transcription?
- Stages of Transcription
- What is Reverse Transcription?
- What is Translation?
- Stages of Translation
- Translation vs Transcription
- Test-Taking Strategies
- Study Annotations
- Join the Study Chat
- Message Not Sent
What is Transcription?
Transcription is the synthesis of an RNA molecule from a DNA template. The enzyme RNA polymerase reads the template strand (also called the antisense strand) of DNA in the 3′ → 5′ direction, building a complementary RNA strand in the 5′ → 3′ direction. The resulting RNA is a single-stranded copy of the gene’s coding information. Three types of RNA can be produced: messenger RNA (mRNA), which carries the protein-coding message; transfer RNA (tRNA), which delivers amino acids during translation; and ribosomal RNA (rRNA), which forms the structural and catalytic core of ribosomes.
A critical base-pairing rule distinguishes RNA from DNA: RNA uses uracil (U) instead of thymine (T). So where DNA pairs A–T and G–C, transcription produces A–U and G–C pairs.
Stages of Transcription
Transcription has four stages: pre-initiation, initiation, elongation, and termination. These differ between prokaryotes and eukaryotes in important ways — prokaryotes transcribe DNA directly in the cytoplasm, while eukaryotes must first uncoil chromatin in the nucleus and perform additional RNA processing steps after transcription.
Pre-Initiation
RNA polymerase (with its σ subunit in prokaryotes) binds to a promoter region upstream of the gene. The promoter is a specific DNA sequence that signals “start here.” Key promoter motifs include TATAAT and TTGACA in prokaryotes, and TATA box (TATAAAA) sequences in eukaryotes. These are cis-acting elements — regulatory sequences on the same DNA molecule as the gene they control. In eukaryotes, transcription factors (proteins) must first bind to the promoter before RNA polymerase can attach — an additional regulatory layer that prokaryotes lack. Once bound, the enzyme denatures (unwinds) the double helix locally, exposing the template strand.
Initiation
RNA polymerase begins synthesizing RNA by inserting the first complementary ribonucleotide at the transcription start site. It then adds subsequent ribonucleotides in the 5′ → 3′ direction, joining them with phosphodiester bonds. At this stage, the growing RNA strand remains hydrogen-bonded to the DNA template.
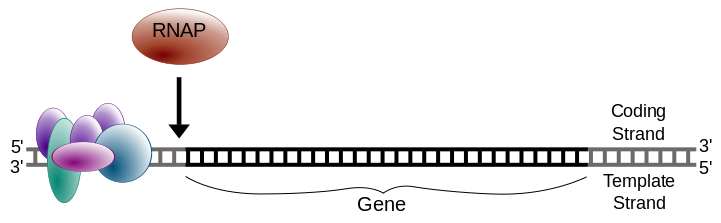
Image Source : Wikipedia
Figure 1: Initiation of transcription. RNAP® refers to RNA polymerase.
Elongation
The σ subunit dissociates, and the core enzyme continues adding ribonucleotides along the template strand. As RNA polymerase advances, the growing RNA strand peels away from the DNA, which re-anneals behind the enzyme. The transcription bubble — the ~17-base-pair region of unwound DNA — moves along the gene.
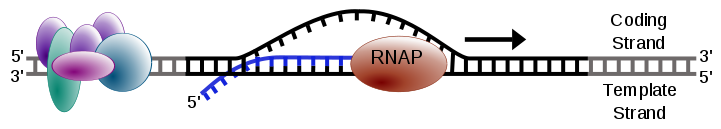
Image Source : Wikipedia
Figure 2: Elongation in transcription
Termination
Transcription ends when RNA polymerase encounters a termination sequence. In prokaryotes, the RNA transcript can form a hairpin loop (a stem-loop structure stabilized by hydrogen bonds) that destabilizes the RNA-DNA hybrid and causes release. Alternatively, the rho (ρ) protein can chase the polymerase and unwind the hybrid. In eukaryotes, termination is coupled with polyadenylation — the addition of a poly-A tail (a string of ~200 adenine nucleotides) to the 3′ end of the transcript.
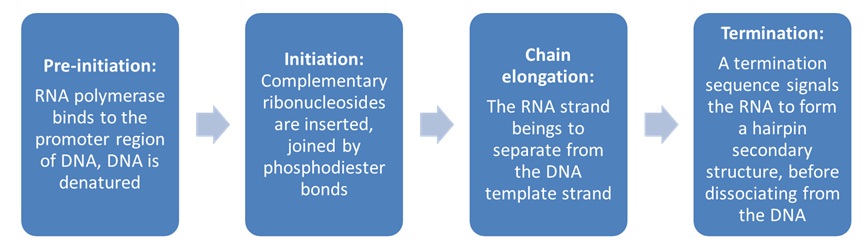
Figure 3: The main events in each stage of transcription
Eukaryotic mRNA processing is an AP® exam favorite. Before the mRNA leaves the nucleus, it undergoes three modifications:
- 5′ cap: A modified guanine nucleotide is added to the 5′ end, protecting the mRNA from degradation and helping ribosomes recognize it.
- Poly-A tail: ~200 adenine nucleotides are added to the 3′ end, stabilizing the mRNA and facilitating nuclear export.
- Splicing: Introns (non-coding sequences) are removed by spliceosomes, and the remaining exons (coding sequences) are joined together. Alternative splicing allows one gene to produce multiple protein variants — a key mechanism of protein diversity.
What is Reverse Transcription?
Reverse transcription is the synthesis of DNA from an RNA template — the reverse of normal transcription. This process is carried out by the enzyme reverse transcriptase and is the hallmark of retroviruses such as HIV. The retrovirus injects its RNA genome into a host cell, reverse transcriptase converts it to double-stranded DNA, and this viral DNA integrates into the host genome using integrase. Once integrated, the host cell’s own machinery transcribes the viral genes, producing new viral particles.
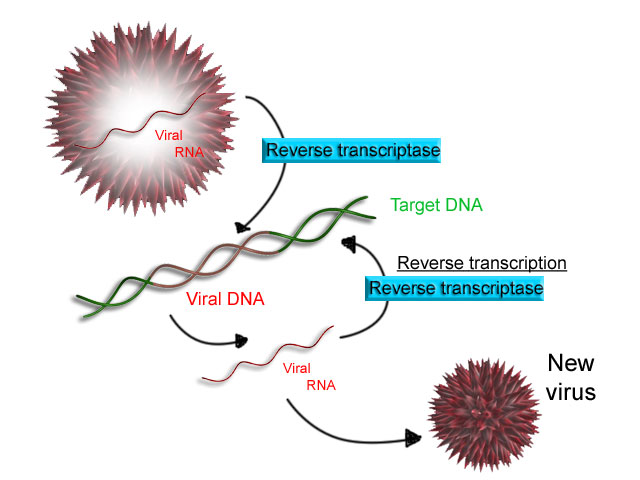
Image Source : Wikipedia
Figure 4: The process of reverse transcription.
Reverse transcription is important for the AP® exam because it demonstrates that the central dogma is not strictly one-directional — information can flow from RNA back to DNA. It also connects to biotechnology: reverse transcriptase is used in the lab to create complementary DNA (cDNA) from mRNA, which is essential for cloning genes and studying gene expression.
What is Translation?
Translation is the process of synthesizing a polypeptide chain from an mRNA template. The ribosome reads the mRNA sequence in sets of three nucleotides called codons, and each codon specifies a particular amino acid (or a stop signal). Transfer RNA (tRNA) molecules serve as adapters — each tRNA carries a specific amino acid and has an anticodon that base-pairs with the complementary mRNA codon.
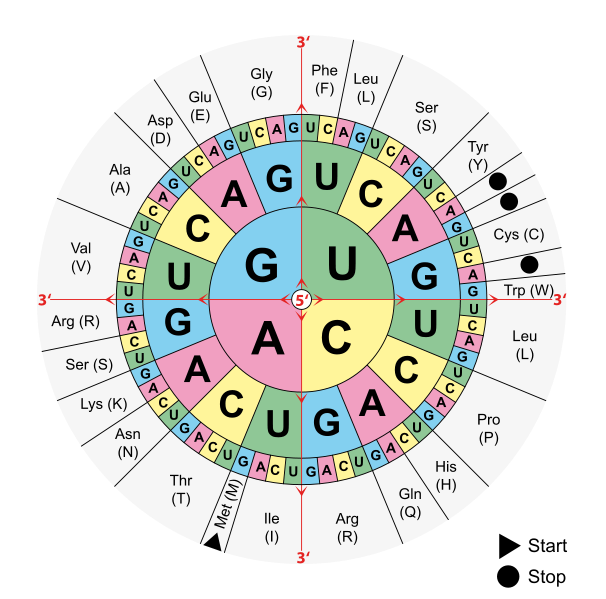
Image Source : Wikimedia Commons
Figure 5: The codon table. Each three-nucleotide codon on mRNA specifies an amino acid. AUG is the start codon (methionine); UAA, UAG, and UGA are stop codons.
The genetic code has several key properties you should know for the AP® exam:
- Universal: Nearly all organisms use the same codon-to-amino-acid assignments — evidence of common ancestry.
- Degenerate (redundant): Multiple codons can specify the same amino acid (e.g., leucine has 6 codons). This redundancy provides a buffer against point mutations.
- Non-overlapping: Codons are read sequentially without sharing nucleotides.
- Unambiguous: Each codon specifies exactly one amino acid (though one amino acid may have multiple codons).
Stages of Translation
Translation occurs in the cytoplasm (prokaryotes) or on ribosomes attached to the rough endoplasmic reticulum (eukaryotes). It proceeds through three stages:
Initiation
The small ribosomal subunit binds to the 5′ end of the mRNA and scans downstream until it encounters the start codon (AUG). An initiator tRNA carrying methionine (Met) base-pairs with AUG at the P site (peptidyl site). Initiation factors (proteins) and GTP hydrolysis facilitate this assembly. The large ribosomal subunit then joins, forming the complete ribosome with three tRNA binding sites: the A site (aminoacyl), P site (peptidyl), and E site (exit).
Elongation
Elongation is a repeating three-step cycle:
- Codon recognition: A charged tRNA with the correct anticodon binds to the codon exposed at the A site. This step requires GTP hydrolysis and elongation factors.
- Peptide bond formation: The enzyme peptidyl transferase (a ribozyme — an RNA with catalytic activity) catalyzes a peptide bond between the amino acid at the P site and the amino acid at the A site. The growing polypeptide chain transfers to the tRNA at the A site.
- Translocation: The ribosome shifts one codon downstream (5′ → 3′). The tRNA at the A site moves to the P site (carrying the polypeptide), the empty tRNA at the P site moves to the E site and exits, and a new codon is exposed at the A site. This step also requires GTP.
This cycle repeats — codon recognition, peptide bond, translocation — adding one amino acid per cycle until the ribosome reaches a stop codon.
Termination
Termination occurs when the ribosome encounters a stop codon (UAA, UAG, or UGA). No tRNA has an anticodon for stop codons. Instead, release factors bind to the A site, triggering hydrolysis of the bond between the polypeptide and the final tRNA. The completed polypeptide is released, and the ribosome disassembles into its subunits.
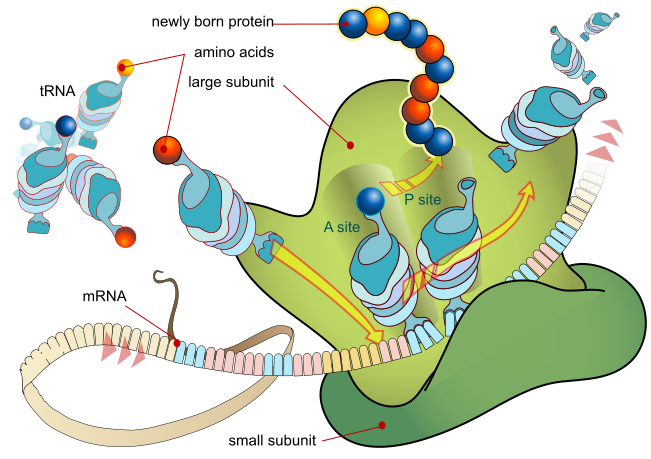
Image Source : Wikipedia
Figure 6: The overview of the process of translation

Figure 7: The main events in each stage of translation.
Translation vs Transcription
The AP® exam frequently asks you to compare these two processes directly. The table below captures the essential distinctions:
| Feature | Transcription | Translation |
|---|---|---|
| Template | DNA (template strand) | mRNA |
| Product | RNA (mRNA, tRNA, or rRNA) | Polypeptide → protein |
| Key enzyme | RNA polymerase | Ribosome (with peptidyl transferase) |
| Direction of synthesis | 5′ → 3′ (reads template 3′ → 5′) | N-terminus → C-terminus (reads mRNA 5′ → 3′) |
| Location (eukaryotes) | Nucleus | Cytoplasm / rough ER |
| Location (prokaryotes) | Cytoplasm | Cytoplasm (can be concurrent with transcription) |
| Monomers used | Ribonucleoside triphosphates (A, U, G, C) | Amino acids (20 types) |
| Bond formed | Phosphodiester bonds | Peptide bonds |
| Start signal | Promoter sequence | Start codon (AUG) |
| Stop signal | Termination sequence / poly-A signal | Stop codon (UAA, UAG, UGA) |
| Energy source | NTP hydrolysis | GTP hydrolysis + charged tRNAs (ATP) |
A key distinction for the AP® exam: in prokaryotes, transcription and translation are coupled — ribosomes begin translating the mRNA while it’s still being transcribed, because there is no nuclear membrane separating the two processes. In eukaryotes, the nuclear envelope enforces a strict separation — transcription occurs in the nucleus, the mRNA is processed (capped, spliced, polyadenylated), exported through nuclear pores, and only then translated in the cytoplasm.
Test-Taking Strategies
Transcription and translation questions appear across Units 6 and 7 of the AP® Biology CED, and they connect to signal transduction, gene regulation, mutations, and biotechnology. Here’s how to handle them on exam day:
- Master the central dogma direction. DNA → RNA → protein. The AP® exam will give you a DNA sequence and ask for the mRNA sequence, or give you mRNA and ask for the amino acid sequence. Practice converting: template DNA 3′-TACGGA-5′ → mRNA 5′-AUGCCU-3′ → Met-Pro. Remember: you read the template strand (not the coding strand) to build mRNA.
- Know where each process occurs. A common MCQ format: “In which cellular compartment does [process] occur?” Transcription = nucleus (eukaryotes) or cytoplasm (prokaryotes). Translation = cytoplasm/rough ER (eukaryotes) or cytoplasm (prokaryotes). Prokaryotic coupling of transcription and translation is a frequent test topic.
- Trace mutation effects through both steps. If an FRQ describes a point mutation in DNA, you need to trace it through transcription (altered mRNA codon) and then translation (altered amino acid — or, if the code is degenerate, possibly no change at all). Silent, missense, nonsense, and frameshift mutations each have distinct consequences — know all four.
- Don’t forget mRNA processing. Eukaryotic mRNA processing (5′ cap, poly-A tail, splicing) is a heavily tested topic. The AP® exam may ask you to explain why a eukaryotic mRNA is shorter than the gene that encoded it (answer: introns were removed). Alternative splicing and its role in protein diversity is another common question.
- Connect to gene regulation. Transcription factors, enhancers, silencers, and promoter accessibility (via chromatin remodeling) control when and how much of a gene is transcribed. These regulatory mechanisms are the foundation of differential gene expression — the reason a liver cell and a neuron have the same DNA but different proteins.
- Use the codon table carefully. The AP® exam provides a codon table. Practice reading it before test day so you don’t waste time. Remember: you look up mRNA codons (not DNA or tRNA sequences) in the table. AUG = start (Met); UAA, UAG, UGA = stop.
- Explain ribosome sites on FRQs. If asked to describe translation, explicitly name the A, P, and E sites and state what happens at each. FRQ rubrics award points for specificity — “tRNA binds to the A site” earns more than “tRNA binds to the ribosome.”
Tag » Where Does Transcription Occur In The Cell
-
Simultaneous Gene Transcription And Translation In Bacteria - Nature
-
Ribosomes, Transcription, Translation | Learn Science At Scitable
-
Where Does Transcription Occur And Where Does ... - Socratic
-
9.3 Transcription – Concepts Of Biology – 1st Canadian Edition
-
Where Does Transcription Occur And Where Does Translation ... - Toppr
-
Steps Of Genetic Transcription | Biology For Majors I - Lumen Learning
-
Stages Of Transcription: Initiation, Elongation & Termination (article)
-
Where Do Transcription And Translation Occur In The Cell? - Quora
-
9.2: Transcription - Biology LibreTexts
-
From DNA To RNA - Molecular Biology Of The Cell - NCBI Bookshelf
-
Definition Of Transcription - NCI Dictionary Of Cancer Terms
-
Where Does Transcription Occur In A Eukaryotic Cell? - Sciencing
-
Protein Synthesis | CK-12 Foundation
-
How Do Genes Direct The Production Of Proteins? - MedlinePlus