1MUH: CRYSTAL STRUCTURE OF TN5 TRANSPOSASE ...
- Careers
- COVID-19
 |
|
- Structure Summary
- Structure
- Annotations
- Experiment
- Sequence
- Genome
- Versions
- FASTA Sequence
- mmCIF Format
- mmCIF Format (Header)
- Legacy PDB Format
- Legacy PDB Format (Header)
- FASTA Sequence
- PDBx/mmCIF Format
- PDBx/mmCIF Format (gz)
- BinaryCIF Format (gz)
- Legacy PDB Format
- Legacy PDB Format (gz)
- PDBML/XML Format (gz)
- Structure Factors (CIF)
- Structure Factors (CIF - gz)
- Validation Full (PDF - gz)
- Validation (XML - gz)
- Validation (CIF - gz)
- Validation 2fo-fc coefficients (CIF - gz)
- Validation fo-fc coefficients (CIF - gz)
- Biological Assembly 1 (CIF - gz)
- Biological Assembly 1 (PDB - gz)
CRYSTAL STRUCTURE OF TN5 TRANSPOSASE COMPLEXED WITH TRANSPOSON END DNA
- PDB DOI: https://doi.org/10.2210/pdb1MUH/pdb
- Entry: 1MUH supersedes: 1F3I
- NAKB: 1MUH
- Classification: TRANSCRIPTION/DNA
- Organism(s): Escherichia coli
- Expression System: Escherichia coli
- Mutation(s): Yes
- Deposited: 2002-09-23 Released: 2002-09-27
Experimental Data Snapshot
- Method: X-RAY DIFFRACTION
- Resolution: 2.30 Å
- R-Value Free: 0.267 (Depositor), 0.200 (DCC)
- R-Value Work: 0.204 (Depositor), 0.200 (DCC)
- R-Value Observed: 0.206 (Depositor)
wwPDB Validation 3D Report Full Report
 This is version 1.5 of the entry. See complete history. LiteratureDownload Primary Citation
This is version 1.5 of the entry. See complete history. LiteratureDownload Primary Citation  Download Mendeley
Download Mendeley
Three-dimensional structure of the Tn5 synaptic complex transposition intermediate.
Davies, D.R., Goryshin, I.Y., Reznikoff, W.S., Rayment, I.(2000) Science 289: 77-85
- PubMed: 10884228 Search on PubMed
- DOI: https://doi.org/10.1126/science.289.5476.77
- Primary Citation of Related Structures: 1MUH
- PubMed Abstract:
Genomic evolution has been profoundly influenced by DNA transposition, a process whereby defined DNA segments move freely about the genome. Transposition is mediated by transposases, and similar events are catalyzed by retroviral integrases such as human immunodeficiency virus-1 (HIV-1) integrase. Understanding how these proteins interact with DNA is central to understanding the molecular basis of transposition. We report the three-dimensional structure of prokaryotic Tn5 transposase complexed with Tn5 transposon end DNA determined to 2.3 angstrom resolution. The molecular assembly is dimeric, where each double-stranded DNA molecule is bound by both protein subunits, orienting the transposon ends into the active sites. This structure provides a molecular framework for understanding many aspects of transposition, including the binding of transposon end DNA by one subunit and cleavage by a second, cleavage of two strands of DNA by a single active site via a hairpin intermediate, and strand transfer into target DNA.
View More Organizational Affiliation: - Department of Biochemistry, University of Wisconsin, Madison, WI 53706, USA.
Explore in 3D: Structure | Sequence Annotations | Electron Density | Validation Report | Ligand Interaction (MN)
Biological Assembly 1Explore in 3D: Structure | Sequence Annotations | Electron Density | Validation Report | Ligand Interaction (MN)
Global Symmetry: Cyclic - C2 (Explore in 3D)Global Stoichiometry: Homo 2-mer - A2 LessFind Similar AssembliesBiological assembly 1 assigned by authors.
PreviousNextMacromolecule Content
- Total Structure Weight: 66.19 kDa
- Atom Count: 4,691
- Modeled Residue Count: 495
- Deposited Residue Count: 521
- Unique protein chains: 1
- Unique nucleic acid chains: 2
- 100%
- 95%
- 90%
- 80%
- 70%
- 60%
- 50%
- 40%
- 30%
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tn5 transposase | C [auth A] | 481 | Escherichia coli | Mutation(s): 3 EC: 3.1 | 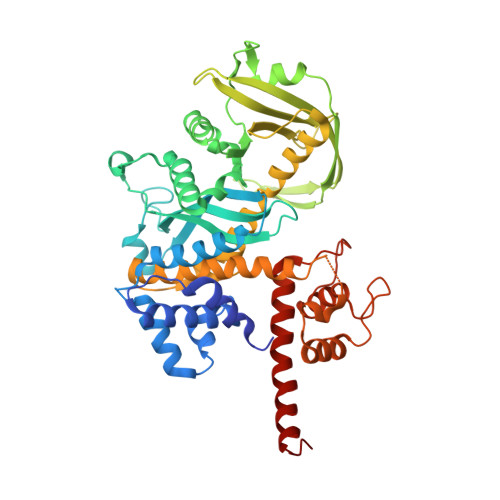 |
UniProt | |||||
| Find proteins for Q46731 (Escherichia coli)Explore Q46731 Go to UniProtKB: Q46731 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q46731 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA TRANSFERRED STRAND | A [auth B] | 20 | N/A | 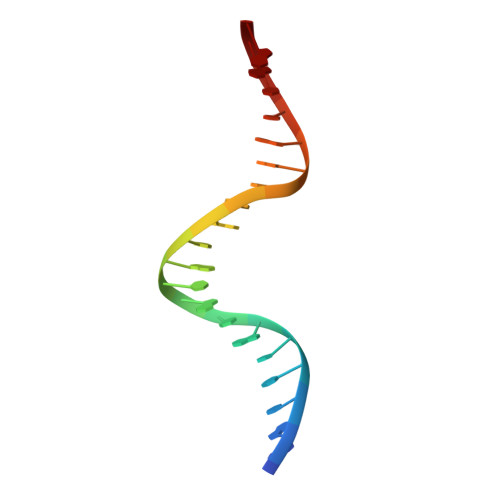 | |
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA NON-TRANSFERRED STRAND | B [auth C] | 20 | N/A | 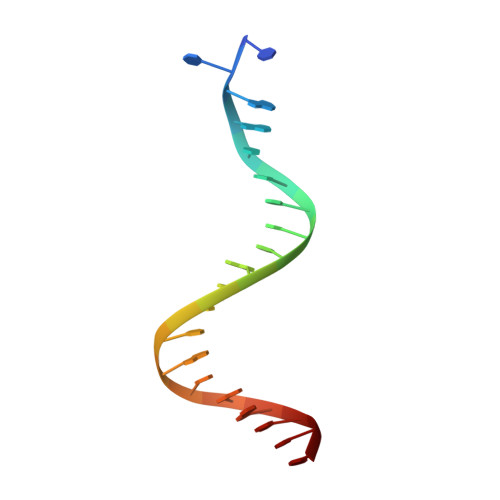 | |
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
MNQuery on MNDownload Ideal Coordinates CCD File Download Instance Coordinates
| D [auth A] | MANGANESE (II) IONMnWAEMQWOKJMHJLA-UHFFFAOYSA-N |  | Interactions
| |
MGQuery on MGDownload Ideal Coordinates CCD File Download Instance Coordinates
| E [auth A] | MAGNESIUM IONMgJLVVSXFLKOJNIY-UHFFFAOYSA-N |  | Interactions
| |
Experimental Data
- Method: X-RAY DIFFRACTION
- Resolution: 2.30 Å
- R-Value Free: 0.267 (Depositor), 0.200 (DCC)
- R-Value Work: 0.204 (Depositor), 0.200 (DCC)
- R-Value Observed: 0.206 (Depositor)
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 113.7 | α = 90 |
| b = 113.7 | β = 90 |
| c = 228.1 | γ = 120 |
| Software Name | Purpose |
|---|---|
| TNT | refinement |
| SCALEPACK | data scaling |
| CNS | refinement |
| DENZO | data reduction |
| CNS | phasing |
Structure Validation
View Full Validation Report
 View more in-depth experimental dataEntry History
View more in-depth experimental dataEntry History Deposition Data
- Released Date: 2002-09-27 Deposition Author(s): Thoden, J.B., Holden, H.M., Davies, D.R., Goryshin, I.Y., Reznikoff, W.S., Rayment, I.
- This entry supersedes: 1F3I
Revision History (Full details and data files)
- Version 1.0: 2002-09-27Type: Initial release
- Version 1.1: 2008-04-28Changes: Version format compliance
- Version 1.2: 2011-07-13Changes: Version format compliance
- Version 1.3: 2019-07-24Changes: Data collection, Refinement description
- Version 1.4: 2021-10-27Changes: Database references, Derived calculations
- Version 1.5: 2024-02-14Changes: Data collection
- About
- About Us
- Citing Us
- Publications
- Team
- Careers
- Usage & Privacy
- Support
- Contact Us
- Help
- Website FAQ
- Glossary
- Service Status
- RCSB PDB is hosted by




- RCSB PDB is a member of
- RCSB Partners
- Nucleic Acid Knowledgebase
- wwPDB Partners
- RCSB PDB
- PDBe
- PDBj
- BMRB
- EMDB


RCSB PDB Core Operations are funded by the U.S. National Science Foundation (DBI-2321666), the US Department of Energy (DE-SC0019749), and the National Cancer Institute, National Institute of Allergy and Infectious Diseases, and National Institute of General Medical Sciences of the National Institutes of Health under grant R01GM157729. RCSB PDB uses resources of the National Energy Research Scientific Computing Center (NERSC), a Department of Energy User Facility.
Từ khóa » Tn5
-
Insertion Site Preference Of Mu, Tn5, And Tn7 Transposons
-
The Tn5 Transposon - PubMed
-
Tn5 Transposase Applied In Genomics Research - PMC - NCBI
-
Tn5 In Vitro Transposition
-
Biochemical Analysis Of Nucleosome Targeting By Tn5 Transposase
-
Tagmentase (Tn5 Transposase) -unloaded - Diagenode
-
Comprehensive Understanding Of Tn5 Insertion Preference Improves ...
-
PTXB1-Tn5 (Plasmid #60240) - Addgene
-
Transposase-assisted Tagmentation Of RNA/DNA Hybrid Duplexes
-
Trans Catalysis In Tn5 Transposition - PNAS
-
Tn5 Transposase And Tagmentation Procedures For Massively Scaled ...
-
[PDF] EZ-Tn5™
Tnp Transposome™ Kit - Lucigen -
Facilitated Large-Scale Sequence Validation Platform Using Tn5 ...








