AirID, A Novel Proximity Biotinylation Enzyme, For Analysis Of Protein ...
TurboID was recently reported as an improved BioID enzyme (Branon et al., 2018). As TurboID has an R118S mutation (RS mutant) that increases the activity of proximity biotinylation, we made RS mutants of the two ancestral BirA enzymes and compared their proximity biotinylation activities in vitro and in cells. An interaction between N-terminal AGIA-BirA-fusion p53 (AGIA-BirA-p53) and FG-MDM2 was used to validate proximity biotinylation ability in vitro. Incubation time, biotinylation temperature, and biotin concentration were investigated as conditions for proximity biotinylation. Consequently, TurboID, AVVA-RG, AVVA-RS, and GFVA-RS showed higher proximity biotinylation activity after 3 hr than did BioID with a 16-hr incubation (Figure 2A). The GFVA enzyme with a RS mutation dramatically increased the activity of proximity biotinylation to RelA (GFVA-RS in Figure 2A and Figure 2—figure supplement 1), but proximity biotinylation was almost the same in AVVA-RG and -RS. AVVA-RG, AVVA-RS, and GFVA-RS showed high activity of proximity biotinylation at temperatures above 16 °C (Figure 2—figure supplement 1A). AVVA-RG, AVVA-RS, and GFVA-RS showed high proximity biotinylation activity at biotin concentrations greater than 0.5 µM (Figure 2—figure supplement 1B). On the basis of these results, three BirA enzymes—AVVA-RG, AVVA-RS, and GFVA-RS—were used for further analysis.
Figure 2 with 5 supplements see all Download asset Open asset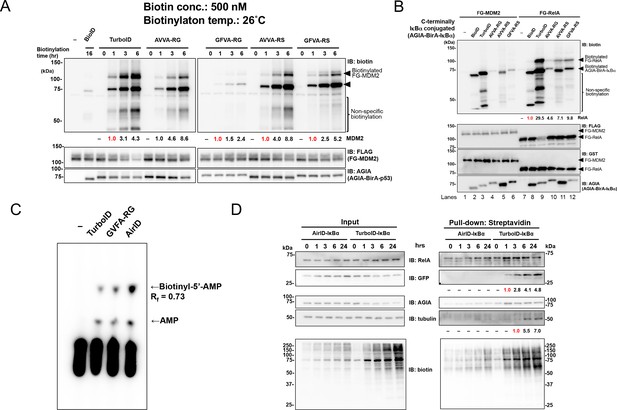
Validation of PPI dependency of novel designed BirA enzymes.
(A) RS mutants of AVVA and GFVA were cloned, and biotinylations of FLAG-GST-MDM2 (FG-MDM2) by BirA-p53 including RS mutants were analyzed. The reaction was performed at 500 nM of biotin at 26 °C for the described time. As a control, the expression levels of both BirA-p53 and MDM2 were detected using anti-AGIA antibody and anti-FLAG antibody, respectively. The band intensity of biotinylated MDM2 was quantified with image J software. The index intensity (value 1.0) is shown in red characters. (B) FG-RelA biotinylation by BirA-IκBα was examined. FG-MDM2 was used as the negative control. Biotinylations were performed at 500 nM of biotin at 26 °C for 1 hr (TurboID), 3 hr (AVVA-RG, AVVA-RS, and GFVA-RS), or 16 hr (BioID). As a control, the expression levels of BirA-p53 and MDM2 were detected using anti-AGIA antibody, anti-FLAG antibody and anti-GST antibody. The band intensity of biotinylated RelA was quantified with image J software. The index intensity (value 1.0) is shown in red characters. (C) GFVA-RG and GFVA-RS expressed using E. coli were purified using Ni beads and mixed with His-bls-FLAG-GST, which was synthesized using a wheat cell-free system and purified using glutathione beads. The mixtures were incubated a solution including [α-32P]ATP and biotin for 30 min at 37 °C. The resultant biotinyl-5′-AMP, AMP, or unreacted ATP was separated using cellulose thin-layer chromatography. (D) GFP and either AirID-IκBα or TurboID-IκBα were transfected in HEK293T, and biotin was added to 5 µM of this mixture for the described time period. After transfecting for 24 hr, cells were lysed by RIPA buffer including protease inhibitors, and biotinylated proteins were pulled down with streptavidin beads. As a control, the expression levels of enzyme-fused protein and target proteins were detected using each protein-specific antibody (left panel). The band intensity of pulled-down GFP and tubulin was quantified with image J software. The index intensity (value 1.0) is shown in red characters.
We used IκBα and RelA to validate the proximity biotinylation ability of these three enzymes in other protein–protein interactions because the IκBα–RelA interaction has been widely observed (Beg et al., 1992; Baeuerle and Baltimore, 1988). As in the analysis of the p53–MDM2 interaction, N-terminal AGIA-BirA-fusion IκBα (AGIA-BirA-IκBα) and FLAG-GST-RelA (FG-RelA) sequences were constructed. FG-MDM2 was used as a negative control. To compare the abilities of the different enzymes directly, the reactions of all enzymes were carried out under the same conditions. After co-incubating AGIA-BirA-IκBα and FG-RelA, AVVA-RS or GFVA-RS, high RelA biotinylation was indicated (Figure 2B). FG-MDM2 biotinylation by AGIA-BirA-IκBα was not observed.
Từ khóa » đạm Enzyme
-
Protein And Enzyme Activity Assays | Thermo Fisher Scientific - SG
-
Action Of Multi-enzyme Complex On Protein Extraction To Obtain A ...
-
Enzyme-catalyzed Protein Crosslinking - PubMed
-
UbiCREST Enzyme Set Protein K-400 - R&D Systems
-
Enzyme And Protein - Home - Karger Publishers
-
Gel Plus Enzyme Protein Gel 2 Oz. - Epicuren Discovery
-
Enzyme Assisted Protein Extraction From Rapeseed, Soybean, And ...
-
Protein, Enzyme, And ELISA Kits | Bio-Rad
-
Protein, Enzyme, And ELISA Kits - Bio-Rad
-
Enzyme Encapsulation By Protein Cages - RSC Publishing
-
Highly Efficient Enzyme Encapsulation In A Protein Nanocage
-
Protein AND Enzyme Gated Supramolecular Disassembly
-
Enzyme-catalyzed Protein Crosslinking | SpringerLink